| The ATLAS Level 2 Trigger |
The e/gamma Analysis Package.
Purpose
The main purpose of the e/gamma Analysis Package is to provide a simple and robust way for calculating the electrons' (including posistrons) and gammas' selection efficiencies for each of the Trigger Levels of the ATLAS Detector. The electrons can come either from single electron simulated/reconstructed events, or from some interesting physics events (signal), such as:
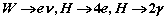
The implementation is extended to include jets, as a background for rate calculations. The package is based on the code written by M. Diaz-Gomez, E. Moyse, V. Perez-Reale, C. Padilla, A. Wildauer. For more information about the original version, visit the initial web-page of the package.
Package location
The package is located at the ATLAS CVS Repository:
offline/Trigger/TrigAnalysis/TrigEgammaAnalysis/
The current, latest version of the package is:
| TrigEgammaAnalysis-00-01-00 |
There is also available the former TrigNtEgamma package. Although this package is not maintained anymore, it is useful for the oldest versions of the trigger e/gamma analysis.
Mailing list - Meetings
The ATLAS mailing list for the package is: atlas-trig-egamma@cern.ch
Subscribers of the list are informed about the regular meetings (normally 1 hour before the PESA meetings) and about the latest modifications and tags of the package.
The agendas and the minutes of the meetings can be found at the CDS Agenda System.
Package description
The following diagram gives the structure of the Package. Click on it to enlarge.
At the Steering File, which has been introduced and replaced the GUI, the user defines the options for running the package.The user can switch between the different data sets without having to build the code again.
Reading the steering file is a case sensitive procedure. Therefore the boolean values should be YES or NO . The other options are:
The histograms are now saved in a file, for the different reconstruction algorithms (i.e Egamma_Histograms_ALGORITHM.root), whenever the user asks for producing them. The histograms are saved in different directories into the file, depending on the level they refer to. Then, it is easy for the user to apply any draw options on the histograms or even to compare histograms from different data sets and different reconstruction algorithms.
Tips:
Production ntuples
The productions refer to single electron (or positron) events with Pt = 25 GeV (Low Luminosity) and with Pt = 30 GeV (High Luminosity). The di-jet events are produced with 17 GeV jets (Low Luminosity) and 25 GeV jets (High Luminosity).
For a further and more detailed description about the production's features, visit the site at RAL
Find below the location of the ntuples.
/castor/cern.ch/atlas/project/dc1/recon/dc1.002026.lumi02.recon.010/dc1.002026.lumi02.recon.010._00001-00010.test.e_minus_25_250.root (1 ntuple with electrons)
/castor/cern.ch/atlas/transfer/emoyse/dc1.002027.lumi02.recon.eplus25.root (1 ntuple with positrons)
/castor/cern.ch/atlas/project/dc1/recon/dc1.002021.lumi10.recon.010/dc1.002021.lumi10.recon.010._00001-00040.test.e_minus_30_250.root (1 ntuple with electrons)
/castor/cern.ch/atlas/transfer/emoyse/dc1.002023.lumi10.recon.eplus30.root (1 ntuple with positrons)
/castor/cern.ch/atlas/project/dc1/recon/dc1.002000.lumi02.recon.010/
/castor/cern.ch/atlas/project/dc1/recon/dc1.002001.lumi10.recon.010/
/castor/cern.ch/user/b/baines/trigprod/rel702++2003-12-12/lumi02/002026/his/ (10 ntuples + 10 ntuples with the KineZ )
/castor/cern.ch/user/b/baines/trigprod/rel702++2003-12-12/lumi10/002021/his/ (40 ntuples + 40 ntuples with the KineZ)
/castor/cern.ch/user/b/baines/trigprod/8.0.2/2004-05-14/lumi02/002026/his/ (10 ntuples)
Running the package
cmt co -r TrigEgammaAnalysis-xx-xx-xx Trigger/TrigAnalysis/TrigEgammaAnalysis
mkdir /home/EgammaAnalysis/
For example: /home/EgammaAnalysis/ProductionNtuples/8.0.2/
tar xzvf TrigEgammaAnalysis.tar.gz
For the above example, you should write: Path8 (Low or High) = /home/EgammaAnalysis/ProductionNtuples/8.0.2/
Tip: A search for TString Path will give you the paths for all the releases used.
make
./main.exe
Tip: As quite often the castor server is down, it is strongly advised to run the package locally. You will only have to transfer the ntuples once. Keep the Analysis.C file so that, for the future package updates, you won't have to fix the paths from the scratch.
Definition of the variables used
| Variable's Name | Description |
| L1em_nroi | Number of EM RoI defined by Level1 |
| L1em_core | |
| L1em_emclus | Energy in the central four cells |
| L1em_emisol | Energy in the EM layer isolation ring |
| L1em_hdisol | Energy deposited in hadronic ring |
| L1em_hdcore | Energy deposited in hadronic core |
| L1em_eta | Eta of the EM cluster defined by LVL1 |
| L1em_phi | Phi of the EM cluster defined by LVL1 |
| Etagen | Generated Eta of the particle |
| Phigen | Generated Phi of the particle |
| Ptgen | Generated Transverse Momentum of the particle |
| Variable's Name | Description |
| T2canclus | Number of LVL2 e/m clusters |
| T2caeta | Weighted eta position of the e/m cluster, calculated in sampling. 2 in a 3x7 cell window |
| T2caphi | Weighted phi position of the e/m cluster, calculated in sampling 2 in a 3x7 cell window |
| T2caeme | Total energy deposited in LAr EM |
| T2cahade | Total Energy deposited in LAr HEC and TileCal |
| T2carcore | E3x7/E7x7 in samp. 2 with Enxm is the energy deposited in a window of nxm cells around the refined LVL1 position |
| T2caeratio | (E1st - E2nd) / (E1st + E2nd) in sampling 1, where E1st and E2nd are the energies of the two highest maxima |
| Variable's Name | Description |
| T2idntracks | Number of Tracks in the Inner Detector |
| T2idalgo | Algorithm that created the track (0:Offline IDScan, 1:SiTrack, 2:Online IDScan, 3:TrtLUT, 4:TrtXk) |
| T2idpt | The transverse momentum of the reconstructed track |
| T2ideta | The eta of the reconstructed track |
| T2idd0 | R of closest approach to origin |
| T2idz0 | Z at closest approach to origin |
| T2idphi0 | Phi at closest approach to origin |
| Variable's Name | Description |
| egnc | Number of reconstructed cluster |
| eget | Transverse Energy of the cluster |
| egf1 | Fraction of energy found in EM Sampling 1 |
| egeta | Eta of the cluster |
| egphi | Phi of the cluster |
| ege277 | Uncorrected energy in 7x7 cells in EM Sampling 2 |
| egisem | e-ID flag. 0: electron, >=1: jet |
| ege237 | Uncorrected energy in 3x7 cells in EM Sampling 2 |
| egetha1 | Et leakage into Sampling 1 of Hadronic Calorimeter |
| egemins1 | Energy of strip with minimum between maximum 1 and 2 |
| ege2tsts1 | 2nd maximum in strips |
| egweta2 | Corrected width in 3x5 cells in the 2nd Sampling |
| egfracs1 | Energy in strips outside core [( (E+-7) - (E+-3) ) / (E+-7)] |
| egwtots1 | Total width in EM Sampling 1 in 20 strips |
| egweta1 | Corrected width in 3 strips in the 1st Sampling |
| et37 |
| Variable's Name | Description |
| egtrkmatchx | link to best matched track in ntuple (for xKalman tracks) |
| Pattern | Bit pattern of SCT and Pixel hits associated |
| Nsihits | Number of SCT and Pixel holes |
| A0vert | Impact Parameter |
| Sharehits | Number of Hits Shared with another reconstructed track |
| Variable's Name | Description |
| egeoverpx | Energy match of cluster with track, E(calo)/p(track) |
| egdeta1x | Delta eta of track extrapolated to Calo S1 |
| egdeta2x | Delta eta of track extrapolated to Calo S2 |
| Nstrawhits | Number of straws associated with the track |
| Ntrhits | Number of Transition Radiation hits associated with the track |
Cuts applied
The cuts which apply at each Trigger Level are of major importance in optimizing the selection efficiencies. It is on the schedule to perform a detail study for optimizing the cuts. Untill then, it is suggested to keep the cuts applied until now. While running the code, one must make sure that the following values HAVE BEEN USED in order for the results to be comparable.
The values-together with the variables (for both luminosities)-can also be found in the /doc directory at the CVS-repository.
The cuts listed below refer to single electron events.
| Low Luminosity | High Luminosity | |
| Level 1 Calo | ||
| ET | > 19 GeV | > 20 GeV |
| EM ring isolation | < 3 GeV | < 5 GeV |
| Hadronic ring isolation | < 2 GeV | < 3 GeV |
| Hadronic core isolation | < 2 GeV | < 2 GeV |
| Level 2 Calo | ||
| ET | > 22.5 GeV | > 25.5 GeV |
| Rηshape | > 0.9 | > 0.9 |
| Rηstrip | > 0.72 | > 0.75 |
| EThad | < 1 GeV | < 2.2 GeV |
| Level 2 ID | ||
| PTtrack | > 8 GeV | > 8 GeV |
| Level 2 ID-Calo | ||
| η-ranges | 0 - 1 , 1 - 1.5 , 1.5 - 2 , 2 - 10 | 0 - 1 , 1 - 1.5 , 1.5 - 2 , 2 - 10 |
| |ΔΦ| | < 0.035, 0.035, 0.03, 0.025 | < 0.05, 0.05, 0.05, 0.05 |
| |Δη| | < 0.07, 0.06, 0.05, 0.05 | < 0.05, 0.05, 0.05, 0.05 |
| ET/PT in the range | [0.2, 3], [0.2, 3], [0.2, 3], [0.2, 3.5] | [0.2, 4], [0.2, 4], [0.2, 5], [0.2, 7] |
| Event Filter Calo | ||
| ET | > 22 GeV | > 27 GeV |
| Event Filter ID | ||
| Number of precision hits | >= 7 | >= 7 |
| Number of pixel hits | >= 1 | >= 1 |
| Number of B-layer hits | >= 1 | >= 1 |
| Impact Parameter | < 0.2 cm | < 0.2 cm |
| Event Filter ID-Calo | ||
| If |η| < 1.37, ET/PT in the range | [0.7, 1.7] | [0.7, 1.7] |
| If |η| >= 1.37, ET/PT in the range | [0.7, 2.7] | [0.7, 2.6] |
| |ΔΦ| | < 0.02 | < 0.02 |
| |Δη| | < 0.01 | < 0.01 |
Results
Useful notes/remarks
