Clatterbridge/TOPAS: Difference between revisions
JacintaYap (talk | contribs) No edit summary |
JacintaYap (talk | contribs) No edit summary |
||
| Line 1: | Line 1: | ||
This simulation is a model of the 62.5 MeV ocular proton beam at the [http://www.clatterbridgecc.nhs.uk/patients/treatment-and-support/proton-therapy Clatterbridge Cancer Centre] built using TOPAS, a wrap around of the Geant4 code specifically developed for proton therapy applications. This model was redeveloped to incorporate geometry updates and general progress made from the previous [http://www.hep.ucl.ac.uk/pbt/wiki/Clatterbridge/Geant4 Geant4 model]. | This simulation is a model of the 62.5 MeV ocular proton beam at the [http://www.clatterbridgecc.nhs.uk/patients/treatment-and-support/proton-therapy Clatterbridge Cancer Centre] built using TOPAS, a wrap around of the Geant4 code specifically developed for proton therapy applications. This model was redeveloped in TOPAS to incorporate geometry updates and general progress made from the previous [http://www.hep.ucl.ac.uk/pbt/wiki/Clatterbridge/Geant4 Geant4 model]. | ||
= Parameter system = | = Parameter system = | ||
| Line 8: | Line 8: | ||
= Geometry = | = Geometry = | ||
In general, TOPAS geometry can be described in different ways: essentially the dimensions and position must be specified with respect to the centre of their parent volume. The component name is included in each line with the stated type (shape), parent volume and material. Translational and rotational ''X, Y, Z'' parameters are assigned for positioning of the component within its mother in ''x'', ''y'' and ''z''. The orientation however will depend on the type of component and additional parameters may be required. | |||
== Treatment beamline == | == Treatment beamline == | ||
| Line 18: | Line 21: | ||
[[File:F_CAD_treatmentline.png]] | [[File:F_CAD_treatmentline.png]] | ||
The geometries were built in CAD according to their exact physical dimensions which are displayed in this [[Media:A_CCC_treatmentline_drawing.pdf|complete schematic]]. Components are exported as Stereolithography binary (STL) files which describe geometry only by their surfaces (a tessellation of triangular and quadrangular faces which approximate the volume). Although other file formats (i.e. GDML) may intrinsically include different properties such as materials or colour attributes, TOPAS does not interpret this information. Therefore each component (with their elements grouped by material) exists as an individual STL file. However, each component may have its own respective axis within the world coordinate system retained during export and additional rotation or translation using parameter lines may be required. To avoid overlaps, a 0.05 mm tolerance was built betwee adjacents surface and components. The positions of every single element within the treatment line coordinate system are displayed in this [[Media:A_CCC_dim_spreadsheet_topas.png|listing]]. | |||
= Physics settings = | = Physics settings = | ||
| Line 35: | Line 41: | ||
= Running the simulation = | = Running the simulation = | ||
== TOPAS == | |||
The simulation has been updated for compatibility with TOPAS v3.6.1 and can be cloned from this [https://github.com/jacyap/ClatterbridgeTreatmentLine.git Github repository]. | |||
= Output = | = Output = | ||
== Analysis == | == Analysis == | ||
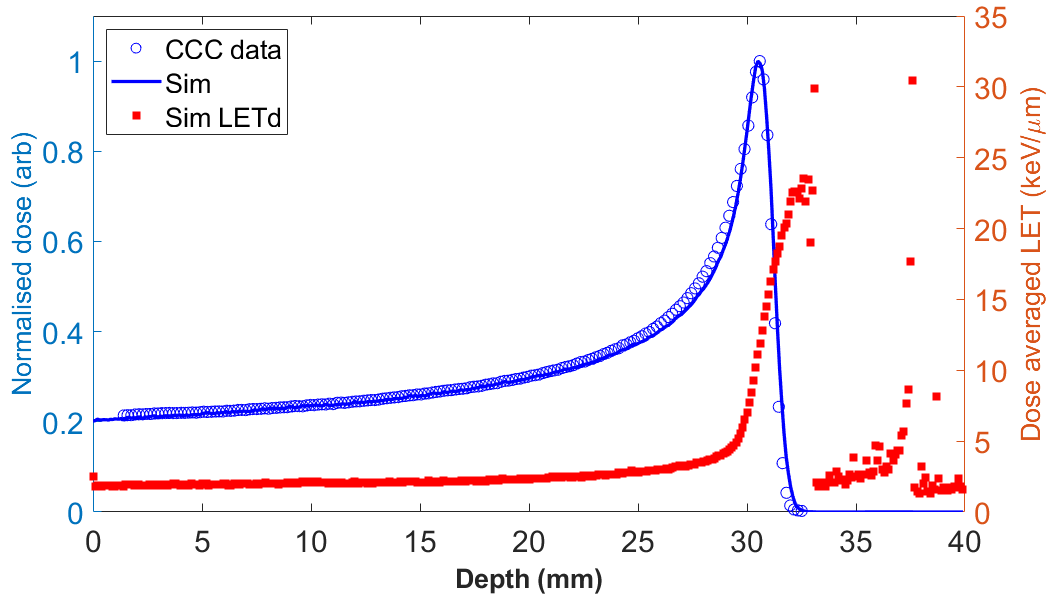
== Dose profiles == | |||
[[File:F_TOPAS_matchBP.png|700px]] | |||
= Additional capabilities = | = Additional capabilities = | ||
Revision as of 04:45, 1 April 2021
This simulation is a model of the 62.5 MeV ocular proton beam at the Clatterbridge Cancer Centre built using TOPAS, a wrap around of the Geant4 code specifically developed for proton therapy applications. This model was redeveloped in TOPAS to incorporate geometry updates and general progress made from the previous Geant4 model.
Parameter system
File chain structure
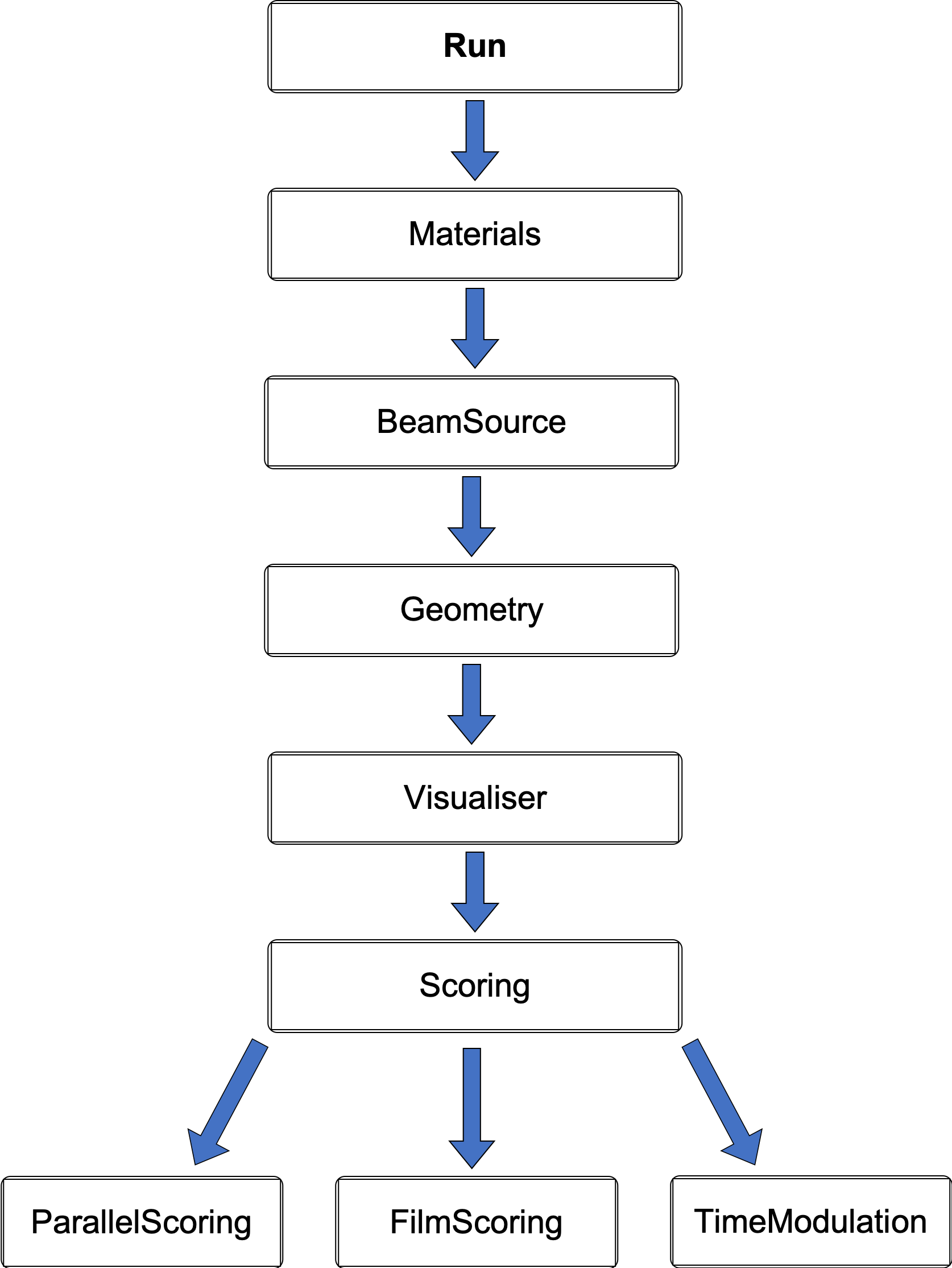
Geometry
In general, TOPAS geometry can be described in different ways: essentially the dimensions and position must be specified with respect to the centre of their parent volume. The component name is included in each line with the stated type (shape), parent volume and material. Translational and rotational X, Y, Z parameters are assigned for positioning of the component within its mother in x, y and z. The orientation however will depend on the type of component and additional parameters may be required.
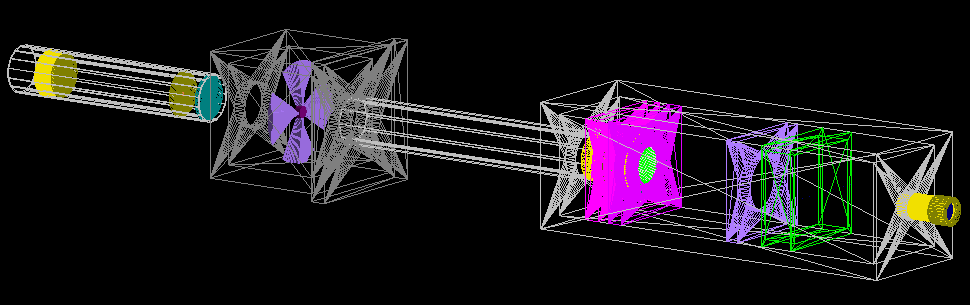
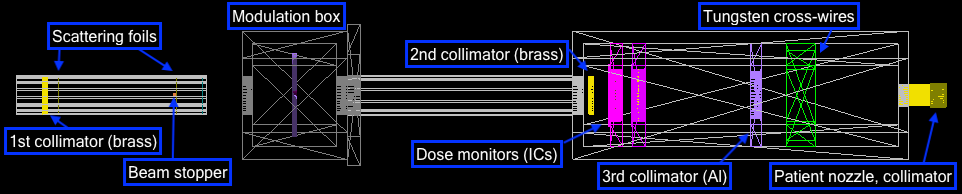
Treatment beamline
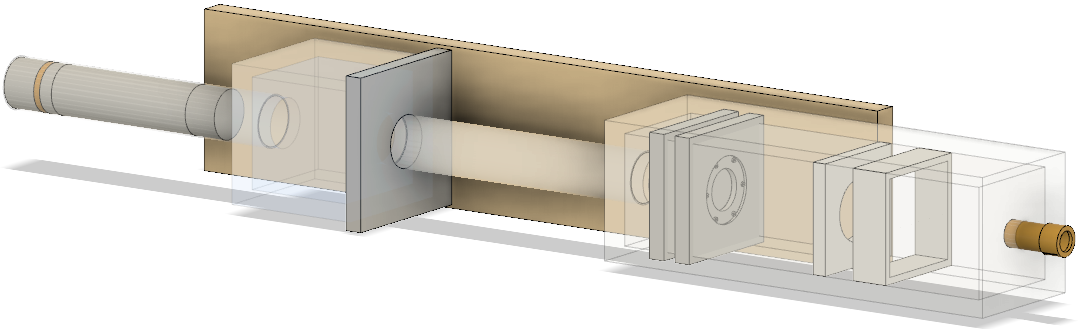
CAD model
The geometries were built in CAD according to their exact physical dimensions which are displayed in this complete schematic. Components are exported as Stereolithography binary (STL) files which describe geometry only by their surfaces (a tessellation of triangular and quadrangular faces which approximate the volume). Although other file formats (i.e. GDML) may intrinsically include different properties such as materials or colour attributes, TOPAS does not interpret this information. Therefore each component (with their elements grouped by material) exists as an individual STL file. However, each component may have its own respective axis within the world coordinate system retained during export and additional rotation or translation using parameter lines may be required. To avoid overlaps, a 0.05 mm tolerance was built betwee adjacents surface and components. The positions of every single element within the treatment line coordinate system are displayed in this listing.
Physics settings
Visualisation
Particle Source
Phase space inputs
BDSIM
Scoring
Volume scorers
Dose deposition
Linear energy transfer
Surface scorers
Transverse beam profiles
Running the simulation
TOPAS
The simulation has been updated for compatibility with TOPAS v3.6.1 and can be cloned from this Github repository.